A number of factors can affect the amount of free Mg 2+ in a reaction, including DNA template concentration, chelating agents in the sample (e.g., EDTA or citrate), dNTP concentration, and the presence of proteins. In contrast, excess free Mg 2+ reduces enzyme fidelity and may increase nonspecific amplification. Without adequate free Mg 2+, PCR polymerases are not active. Magnesium is a required cofactor for thermostable DNA polymerases and is important for successful amplification. What are the critical factors for amplification of long genomic targets? For samples containing excess template (up to 1 µg), we recommend PrimeSTAR GXL DNA Polymerase. It is important to note that not all polymerases can tolerate excessive amounts of template. The amount of cDNA in a PCR reaction can be as little as 10 pg (RNA equivalent). cDNA input is typically described in terms of equivalent RNA input. The amount of cDNA template depends on the copy number of the target.Typically, 1 µg of lambda DNA contains 1.9 x 10 10 molecules of DNA therefore, the template input can be as little as 100 pg.coli genomic DNA contains 2 x 10 8 molecules of DNA therefore, the recommended amount of template is between 100 pg and 1 ng. Template amounts for higher-complexity templates range between 10 ng and 500 ng. High-copy targets, such as housekeeping genes, require only 10 ng of template. For most PCR applications, 30–100 ng of human genomic DNA is sufficient. Typically, 1 µg of human genomic DNA contains 3.04 x 10 5 molecules of DNA.Approximately 10 4 copies of the target DNA sequence are required to detect the amplification product in 25–30 PCR cycles. The optimal amount of template required depends on the complexity of the template and the copy number of the target sequence. 
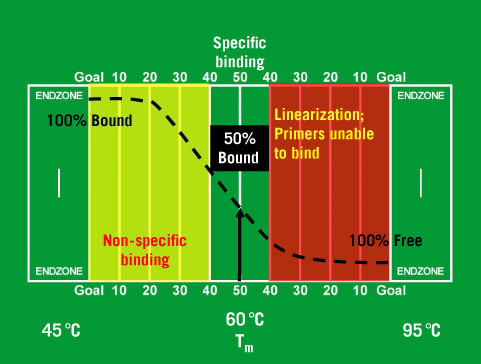
Should I use a three-step or a two-step PCR protocol? If using these enzymes with samples containing excess template, an elongation time of 1 min/kb should be used. PrimeSTAR Max DNA Polymerase and PrimeSTAR GXL DNA Polymerase contain a proprietary elongation factor and allow for high-speed reactions at 5–20 sec/kb. When using the high-speed enzymes SpeedSTAR HS DNA Polymerase or SapphireAmp Fast PCR Master Mix, use a reaction rate of 10 sec/kb of amplified product (i.e., 10 sec for a 1-kb product, 20 sec for a 2-kb product, etc.). In general, an extension time of 1 min/kb is recommended. For GC-rich targets or amplifications of long sequences (>10 kb), a two-step PCR protocol is recommended. When amplifying short sequences smaller than 1 kb, a three-step PCR protocol is recommended.Excessively long annealing times may lead to mispriming-induced nonspecific amplification. Therefore, it is important to use a short annealing time of 5–15 sec. Enzymes in the PrimeSTAR series have excellent priming efficiency.For Taq enzymes, the recommended annealing time is 30 sec.Using annealing temperatures that are too low may result in mispriming and nonspecific amplification, leading to low yields of the desired product.Īmplification efficiency and specificity can be improved by adjusting the annealing temperature according to the primer's T m or by performing two-step PCR. The annealing step should be adjusted for each primer set the annealing temperature depends directly on the T m of primers. If using a heat-resistant enzyme, such as one of the PrimeSTAR polymerases, we recommend a denaturation step of short duration and high temperature (i.e., 5–10 sec at 98☌).ĭenaturation at an excessively high temperature or for too long may result in loss of enzyme activity and/or damage to long templates. A general guideline is 94–95☌ for 30 sec or 98☌ for 10 sec. PrimeSTAR enzymes do not require preheating for enzyme activation.ĭenaturing conditions should be selected by considering the thermal cycler model that will be used.For Takara LA Taq DNA polymerases and Advantage GC2 DNA polymerases, an initial denaturation step is required.For Terra PCR Direct Polymerase Mix, which is used for direct PCR amplification from tissue without DNA extraction and purification, preheating at 98☌ for 2 min is required.Excessive heat treatment may lead to enzyme inactivation. 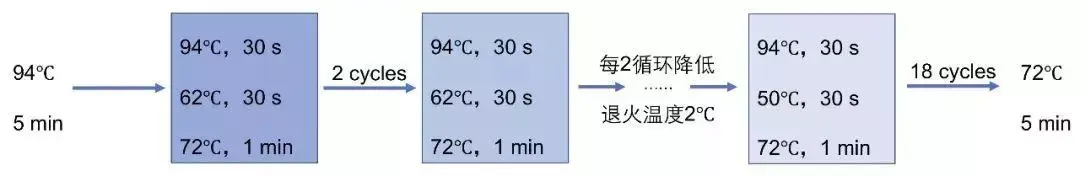
Preheating is sometimes required to denature complex templates (e.g., genomic DNA) 94☌ for 1 min is sufficient for denaturation. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
